It’s time for part 4 of the BBC Good Food Series. In this post we’ll learn how to use the Jaccard Similarity Algorithm to compute recipe to recipe similarities, and more.
You can read the previous parts of the series to get up to date:
Part 1: Importing BBC goodfood information into Neo4j
Part 2: What can I make with these ingredients?
Part 3: A segue into graph modellingSimilarity Algorithms in Neo4j Graph Algorithms Library
In this post we’re going to use the similarity algorithms in the Neo4j Graph Algorithms, but let’s first remind ourselves about the graph model that we created in the earlier posts.
We can view the model by running CALL db.schema()in the Neo4j Browser:

Recipes are at the centre of our world, and they’re connected to Collections, Keywords, Authors, DietTypes, and Ingredients. We could compute similarities between any of this items, but why would we want to do that?
Why compute similarities?
We want to compute similarities between items so that we can make recommendations to users about other things that they might like.
- If a user likes a recipe, can we suggest other recipes that they could try?
Recipe to Recipe
Recipes sit at the centre of this graph, so that seems like a good place to start. We can compute the similarity of a pair of recipes based on the ingredients that they have in common.
We can use the Jaccard Similarity Algorithm. Jaccard Similarity is defined as the size of the intersection divided by the size of the union of two sets, best seen in the diagram below:
Jaccard SimilarityThis algorithm returns a score between 0 and 1 that indicates similarity between items. 0 means that they are dissimilar. 1 means that they are identical.
Let’s have a look how it works with an example. The diagram below shows two bolognese recipes and the ingredients that they contain:
Ingredients used in Bolognese RecipesWe want to compute the similarity of the recipes based on their ingredients. From visual inspection we can see that there are 4 overlapping ingredients: spaghetti, carrot, basil, and olive oil. The combined number of ingredients is 23, which means our Jaccard Similarity score should be 4/23 = 0.17
Jaccard Similarity Function
Let’s use the Jaccard Similarity function to compute the similarity of these two recipes and check that it matches our quick calculation:
// Find the 1st recipe and collect the node ids into a list
MATCH (r1:Recipe)-[:CONTAINS_INGREDIENT]->(i)
WHERE r1.name = "So-simple spaghetti Bolognese"
WITH r1, collect(id(i)) AS r1Ingredients
// Find the 2nd recipe and collect the node ids into a list
MATCH (r2:Recipe)-[:CONTAINS_INGREDIENT]->(i)
WHERE r1.name = "The best spaghetti Bolognese recipe"
WITH r1, r1Ingredients, r2, collect(id(i)) AS r2Ingredients
// Compute similarity based on the lists of node ids
RETURN algo.similarity.jaccard(
r1Ingredients, r2Ingredients) AS score
Cool! The result is as we expected, and the similarity function is great for computing the similarity of pairs of nodes. But what if we want to compute the similarity between all pairs of recipes?
Jaccard Similarity Procedure
It will take a long time to do this one function call at a time, which is where the similarity procedure comes into play. The procedure parallelises the computation of similarities, and is therefore well suited for computing the similarity of lots of pairs of items.
The library contains two procedures for Jaccard Similarity: a streaming procedure and a write procedure.
Streaming procedure
The following query will compute the similarity of all pairs of recipes based on their ingredients, and stream the top 20 pairs of most similar recipes:
// Construct lists of node ids of ingredients,
// grouped by node ids of recipes
MATCH (r:Recipe)-[:CONTAINS_INGREDIENT]->(ingredient)
WITH {item:id(r), categories: collect(id(ingredient))} as userData
WITH collect(userData) as data
CALL algo.similarity.jaccard.stream(data)
YIELD item1, item2, count1, count2, intersection, similarity
// Look up nodes by node id
WITH algo.asNode(item1) AS from,
algo.asNode(item2) AS to,
similarity, intersection
RETURN from.name, from.id, to.name, to.id, intersection, similaritySimilar recipes
ORDER BY similarity DESC
LIMIT 20
All the pairs here have identical ingredients, which in several cases looks like it’s because they are duplicates! Getting rid of those duplicates is an interesting problem, but we’ll leave that for another blog post.
Write procedure
This query only shows us the top 20 similar pairs of recipes. It’d be good to know the distribution of similarity scores, which we can do by calling the write version of the algorithm. We’ll set write:false so that we don’t write any relationships to the database.
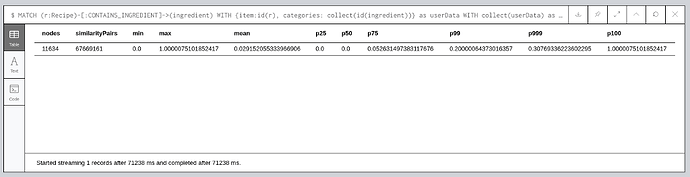
The following query returns the number of similarity pairs, along with 25th, 50th, 75th, 99th, 99.9th, and 100th percentile values:
MATCH (r:Recipe)-[:CONTAINS_INGREDIENT]->(ingredient)Jaccard Similarity Percentiles
WITH {item:id(r), categories: collect(id(ingredient))} as userData
WITH collect(userData) as data
CALL algo.similarity.jaccard(data, {write: false})
YIELD nodes, similarityPairs, min, max, mean,
p25, p50, p75, p99, p999, p100
RETURN nodes, similarityPairs, min, max, mean,
p25, p50, p75, p99, p999, p100
From this output we learn that 50% of the similarity scores being returned are 0, and 99% of them are less than 0.2. We can get rid of those pairs of items by passing in the similarityCutoff config parameter.
Filtering with similarityCutoff
Let’s re-run the write version of the procedure, but this time we’ll return pairs of recipes that have a Jaccard Similarity of 0.2 or higher:
MATCH (r:Recipe)-[:CONTAINS_INGREDIENT]->(ingredient)Jaccard Similarities above 0.2
WITH {item:id(r), categories: collect(id(ingredient))} as userData
WITH collect(userData) as data
CALL algo.similarity.jaccard(data, {
write: false, similarityCutoff: 0.2
})
YIELD nodes, similarityPairs, min, max, mean,
p25, p50, p75, p99, p999, p100
RETURN nodes, similarityPairs, min, max, mean,
p25, p50, p75, p99, p999, p100
The number of similarity pairs has reduced from 676 million to 762,000. We have 11,000 recipes nodes, which is an average of 70 similar values per recipe. For our recipe recommendation engine we won’t show that many similar recipes, a maximum of 10 similar recipes is sufficient. similarityCutoff: 0.3 returns 89,600 similarity pairs, an average of 8 similar values per recipe, so we’ll use that cut off value.
topK
The similarity algorithms all support the topK config parameter. If we set this parameter the procedure will return a maximum of k similar values per node.
We’ll set topK: 5 , which will find the 5 most similar values per recipe. We’ll also set write:true so that relationships will be created between for each of the similar values. The following query will do this:
MATCH (r:Recipe)-[:CONTAINS_INGREDIENT]->(ingredient)
WITH {item:id(r), categories: collect(id(ingredient))} as userData
WITH collect(userData) as data
CALL algo.similarity.jaccard(data, {
write: true,
writeRelationshipType: "SIMILAR",
writeProperty: "score",
similarityCutoff: 0.3
})
YIELD nodes, similarityPairs, min, max, mean,
p25, p50, p75, p99, p999, p100
RETURN nodes, similarityPairs, min, max, mean,
p25, p50, p75, p99, p999, p100
Exploring the similarity graph
Now let’s have a look at the similarity graph that’s been created by running this procedure:
Similarity GraphIf you like potatoes, there are lots of recipes to keep you happy! And let’s see if we can find some other recipes for our bolognese loving friends:
MATCH path = (r1:Recipe)-[:SIMILAR]->(r2)-[:CONTAINS_INGREDIENT]-(i)Other bolognese recipes
WHERE r1.name = "So-simple spaghetti Bolognese"
RETURN path
We could use the results of this query as recommendations for users who want to try a different bolognese recipe.
So we’ve now learnt how to compare recipes against recipes, but what if we want to compare items that have different labels?
Collection to Ingredient
An example of this is computing similarities between collections and ingredients. These types of nodes intersect on recipes, via the following graph pattern:
(collection)<-[:COLLECTION]-(rec)-[:HAS_INGREDIENT]->(ingredient)
We can can build up lists of the node ids of recipes for all collection and ingredient nodes and compare them to each other. If we used the approach we’ve seen so far, we’d have to compare all ingredients and collections to each other, which isn’t what we want. We want to compare collections to ingredients, not collections to collections or ingredients to ingredients.
The similarity algorithms allows us to pass in the sourceIds and targetIds config parameters to handle this.
The following query computes Jaccard similarity between Collection and Ingredients, and returns the top 100 most similar pairs:
// Create list of nodes ids of recipes for collections
MATCH (recipe:Recipe)-[:COLLECTION]->(collection)
WITH {item:id(collection), categories: collect(id(recipe))} as data
WITH collect(data) AS collectionRecipes
// Create list of nodes ids of recipes for ingredients
MATCH (recipe:Recipe)-[:CONTAINS_INGREDIENT]->(ingredient)
WITH collectionRecipes,
{item:id(ingredient), categories: collect(id(recipe))} as data
WITH collectionRecipes, collect(data) AS ingredientRecipes
// Extract node ids of collections to use as sourceIds
// Extract node ids of recipes to use as targetids
WITH collectionRecipes + ingredientRecipes AS allRecipes,
[value in collectionRecipes | value.item] AS sourceIds,
[value in ingredientRecipes | value.item] AS targetIds
CALL algo.similarity.jaccard.stream(allRecipes, {
similarityCutoff: 0.0,
sourceIds: sourceIds,
targetIds: targetIds
})
YIELD item1, item2, count1, count2, intersection, similarity
WITH algo.getNodeById(item1) AS from,
algo.getNodeById(item2) AS to,
similarity
RETURN labels(from) AS fromLabels, from.name AS from,
labels(to) AS toLabels, to.name AS to, similarity
ORDER BY similarity DESC
LIMIT 100Collection -> Ingredient SimilaritiesUnsurprisingly, ingredients with the same name as the collection have high similarity scores.
We can also use this approach to compare a single collection to all ingredients. The query below finds the top 5 ingredients (topK:5) used by the ‘Apple’ collection:
MATCH (recipe:Recipe)-[:COLLECTION]->(collection:Collection)
WHERE collection.name = "Apple"
WITH {item:id(collection), categories: collect(id(recipe))} as data
WITH collect(data) AS collectionRecipes
MATCH (recipe:Recipe)-[:CONTAINS_INGREDIENT]->(ingredient)
WITH collectionRecipes,
{item:id(ingredient), categories: collect(id(recipe))} as data
WITH collectionRecipes, collect(data) AS ingredientRecipes
WITH collectionRecipes + ingredientRecipes AS allRecipes,
[value in collectionRecipes | value.item] AS sourceIds,
[value in ingredientRecipes | value.item] AS targetIds
CALL algo.similarity.jaccard.stream(allRecipes, {
similarityCutoff: 0.0,
sourceIds: sourceIds,
targetIds: targetIds,
topK: 5
})
YIELD item2, intersection, similarity
WITH algo.getNodeById(item2) AS to, similarity
RETURN to.name AS to, similarity
ORDER BY similarity DESCIngredients similar to the Apple CollectionSummary
In this post we’ve learnt how to use the Jaccard Similarity Algorithm in the Neo4j Graph Algorithms Library, and we’ve explored what we can do with the various config parameters that it supports.
If you enjoyed learning how to apply graph algorithms to make sense of data, you might like the O’Reilly Graph Algorithms Book that Amy Hodler and I wrote.
You can download a free copy from neo4j.com/graph-algorithms-book

Download the O’Reilly Graph Algorithms Book
![]()
What’s cooking? Part 4: Similarities was originally published in neo4j on Medium, where people are continuing the conversation by highlighting and responding to this story.